Arsenic »
PDB 2im3-2xnq »
2wta »
Arsenic in PDB 2wta: Acinetobacter Baumanii Nicotinamidase Pyrazinamidease
Enzymatic activity of Acinetobacter Baumanii Nicotinamidase Pyrazinamidease
All present enzymatic activity of Acinetobacter Baumanii Nicotinamidase Pyrazinamidease:
3.5.1.19;
3.5.1.19;
Protein crystallography data
The structure of Acinetobacter Baumanii Nicotinamidase Pyrazinamidease, PDB code: 2wta
was solved by
P.K.Fyfe,
V.A.Rao,
A.Zemla,
S.Cameron,
W.N.Hunter,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 42.54 / 1.70 |
| Space group | I 1 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 50.688, 46.507, 107.602, 90.00, 101.97, 90.00 |
| R / Rfree (%) | 14.748 / 17.661 |
Other elements in 2wta:
The structure of Acinetobacter Baumanii Nicotinamidase Pyrazinamidease also contains other interesting chemical elements:
| Chlorine | (Cl) | 3 atoms |
| Zinc | (Zn) | 1 atom |
Arsenic Binding Sites:
The binding sites of Arsenic atom in the Acinetobacter Baumanii Nicotinamidase Pyrazinamidease
(pdb code 2wta). This binding sites where shown within
5.0 Angstroms radius around Arsenic atom.
In total only one binding site of Arsenic was determined in the Acinetobacter Baumanii Nicotinamidase Pyrazinamidease, PDB code: 2wta:
In total only one binding site of Arsenic was determined in the Acinetobacter Baumanii Nicotinamidase Pyrazinamidease, PDB code: 2wta:
Arsenic binding site 1 out of 1 in 2wta
Go back to
Arsenic binding site 1 out
of 1 in the Acinetobacter Baumanii Nicotinamidase Pyrazinamidease
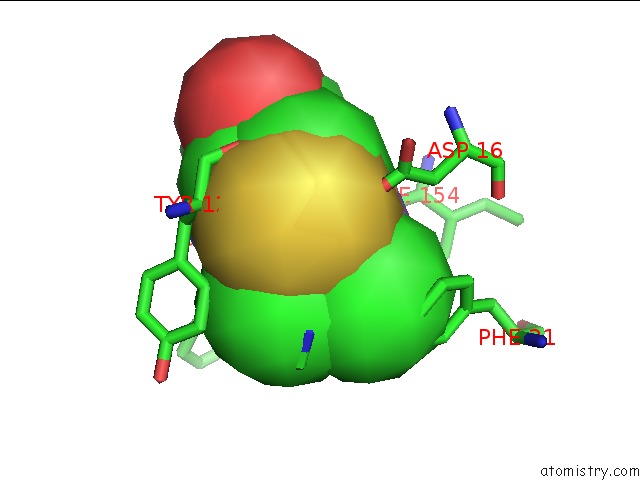
Mono view
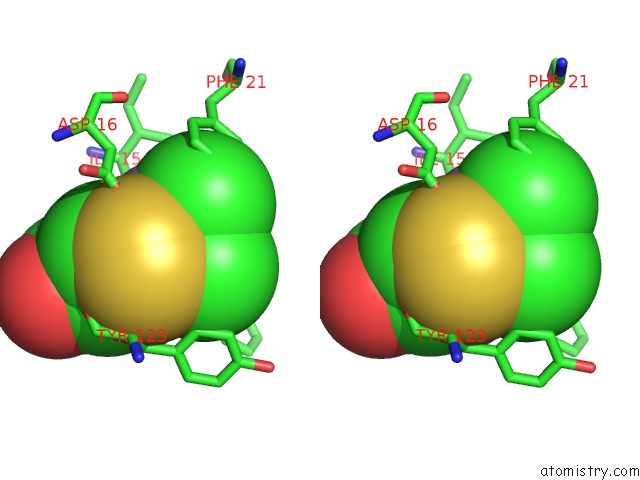
Stereo pair view

Mono view

Stereo pair view
A full contact list of Arsenic with other atoms in the As binding
site number 1 of Acinetobacter Baumanii Nicotinamidase Pyrazinamidease within 5.0Å range:
|
Reference:
P.K.Fyfe,
V.A.Rao,
A.Zemla,
S.Cameron,
W.N.Hunter.
Specificity and Mechanism of Acinetobacter Baumanii Nicotinamidase: Implications For Activation of the Front-Line Tuberculosis Drug Pyrazinamide. Angew.Chem.Int.Ed.Engl. V. 48 9176 2009.
ISSN: ISSN 1433-7851
PubMed: 19859929
DOI: 10.1002/ANIE.200903407
Page generated: Sun Jul 6 23:17:11 2025
ISSN: ISSN 1433-7851
PubMed: 19859929
DOI: 10.1002/ANIE.200903407
Last articles
F in 7K15F in 7K12
F in 7K0W
F in 7JYR
F in 7JY3
F in 7JYQ
F in 7JX9
F in 7JY1
F in 7JY0
F in 7JXW