Arsenic »
PDB 3g3s-3n5t »
3hbz »
Arsenic in PDB 3hbz: Crystal Structure of A Putative Glycoside Hydrolase (BT_2081) From Bacteroides Thetaiotaomicron Vpi-5482 at 2.05 A Resolution
Protein crystallography data
The structure of Crystal Structure of A Putative Glycoside Hydrolase (BT_2081) From Bacteroides Thetaiotaomicron Vpi-5482 at 2.05 A Resolution, PDB code: 3hbz
was solved by
Joint Center For Structural Genomics (Jcsg),
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 29.75 / 2.05 |
| Space group | P 32 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 94.551, 94.551, 107.812, 90.00, 90.00, 120.00 |
| R / Rfree (%) | 15.9 / 19.1 |
Other elements in 3hbz:
The structure of Crystal Structure of A Putative Glycoside Hydrolase (BT_2081) From Bacteroides Thetaiotaomicron Vpi-5482 at 2.05 A Resolution also contains other interesting chemical elements:
| Calcium | (Ca) | 2 atoms |
| Sodium | (Na) | 2 atoms |
Arsenic Binding Sites:
The binding sites of Arsenic atom in the Crystal Structure of A Putative Glycoside Hydrolase (BT_2081) From Bacteroides Thetaiotaomicron Vpi-5482 at 2.05 A Resolution
(pdb code 3hbz). This binding sites where shown within
5.0 Angstroms radius around Arsenic atom.
In total 2 binding sites of Arsenic where determined in the Crystal Structure of A Putative Glycoside Hydrolase (BT_2081) From Bacteroides Thetaiotaomicron Vpi-5482 at 2.05 A Resolution, PDB code: 3hbz:
Jump to Arsenic binding site number: 1; 2;
In total 2 binding sites of Arsenic where determined in the Crystal Structure of A Putative Glycoside Hydrolase (BT_2081) From Bacteroides Thetaiotaomicron Vpi-5482 at 2.05 A Resolution, PDB code: 3hbz:
Jump to Arsenic binding site number: 1; 2;
Arsenic binding site 1 out of 2 in 3hbz
Go back to
Arsenic binding site 1 out
of 2 in the Crystal Structure of A Putative Glycoside Hydrolase (BT_2081) From Bacteroides Thetaiotaomicron Vpi-5482 at 2.05 A Resolution
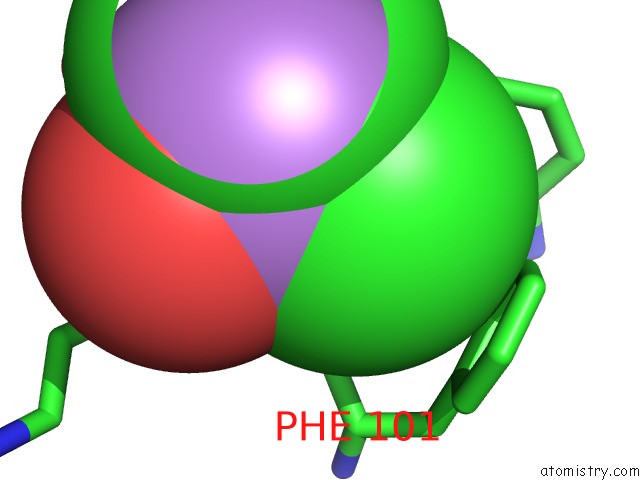
Mono view
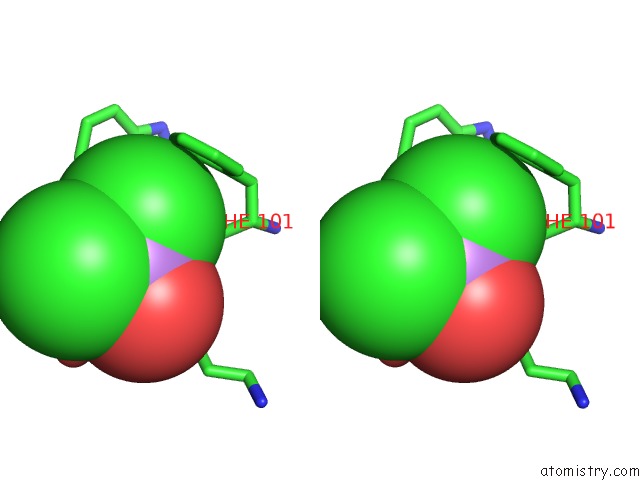
Stereo pair view

Mono view

Stereo pair view
A full contact list of Arsenic with other atoms in the As binding
site number 1 of Crystal Structure of A Putative Glycoside Hydrolase (BT_2081) From Bacteroides Thetaiotaomicron Vpi-5482 at 2.05 A Resolution within 5.0Å range:
|
Arsenic binding site 2 out of 2 in 3hbz
Go back to
Arsenic binding site 2 out
of 2 in the Crystal Structure of A Putative Glycoside Hydrolase (BT_2081) From Bacteroides Thetaiotaomicron Vpi-5482 at 2.05 A Resolution
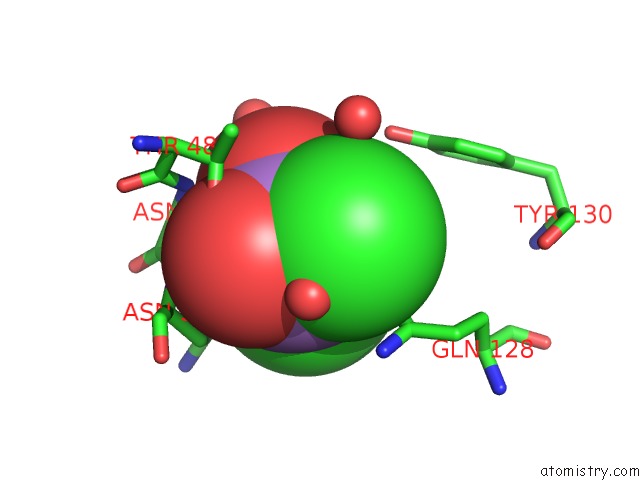
Mono view
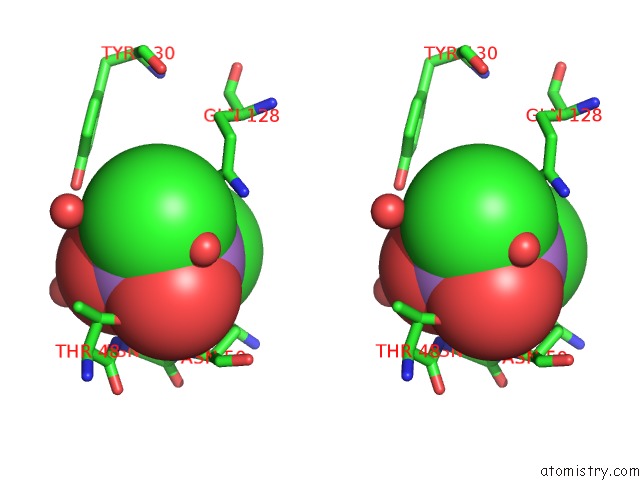
Stereo pair view

Mono view

Stereo pair view
A full contact list of Arsenic with other atoms in the As binding
site number 2 of Crystal Structure of A Putative Glycoside Hydrolase (BT_2081) From Bacteroides Thetaiotaomicron Vpi-5482 at 2.05 A Resolution within 5.0Å range:
|
Reference:
A.P.Yeh,
P.Abdubek,
T.Astakhova,
H.L.Axelrod,
C.Bakolitsa,
X.Cai,
D.Carlton,
C.Chen,
H.J.Chiu,
M.Chiu,
T.Clayton,
D.Das,
M.C.Deller,
L.Duan,
K.Ellrott,
C.L.Farr,
J.Feuerhelm,
J.C.Grant,
A.Grzechnik,
G.W.Han,
L.Jaroszewski,
K.K.Jin,
H.E.Klock,
M.W.Knuth,
P.Kozbial,
S.S.Krishna,
A.Kumar,
W.W.Lam,
D.Marciano,
D.Mcmullan,
M.D.Miller,
A.T.Morse,
E.Nigoghossian,
A.Nopakun,
L.Okach,
C.Puckett,
R.Reyes,
H.J.Tien,
C.B.Trame,
H.Van Den Bedem,
D.Weekes,
T.Wooten,
Q.Xu,
K.O.Hodgson,
J.Wooley,
M.A.Elsliger,
A.M.Deacon,
A.Godzik,
S.A.Lesley,
I.A.Wilson.
Structure of Bacteroides Thetaiotaomicron BT2081 at 2.05 A Resolution: the First Structural Representative of A New Protein Family That May Play A Role in Carbohydrate Metabolism. Acta Crystallogr.,Sect.F V. 66 1287 2010.
ISSN: ESSN 1744-3091
PubMed: 20944224
DOI: 10.1107/S1744309110028228
Page generated: Sun Jul 6 23:24:28 2025
ISSN: ESSN 1744-3091
PubMed: 20944224
DOI: 10.1107/S1744309110028228
Last articles
F in 7GJPF in 7GJO
F in 7GIU
F in 7GIH
F in 7GIL
F in 7GJ4
F in 7GI9
F in 7GIF
F in 7GIB
F in 7GHN