Arsenic »
PDB 3n67-3smt »
3pd8 »
Arsenic in PDB 3pd8: X-Ray Structure of the Ligand-Binding Core of GLUA2 in Complex with (S)-7-Hpca at 2.5 A Resolution
Protein crystallography data
The structure of X-Ray Structure of the Ligand-Binding Core of GLUA2 in Complex with (S)-7-Hpca at 2.5 A Resolution, PDB code: 3pd8
was solved by
K.Frydenvang,
J.S.Kastrup,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 29.32 / 2.48 |
| Space group | P 21 21 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 114.618, 164.002, 47.620, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 17.5 / 26.1 |
Other elements in 3pd8:
The structure of X-Ray Structure of the Ligand-Binding Core of GLUA2 in Complex with (S)-7-Hpca at 2.5 A Resolution also contains other interesting chemical elements:
| Zinc | (Zn) | 5 atoms |
Arsenic Binding Sites:
The binding sites of Arsenic atom in the X-Ray Structure of the Ligand-Binding Core of GLUA2 in Complex with (S)-7-Hpca at 2.5 A Resolution
(pdb code 3pd8). This binding sites where shown within
5.0 Angstroms radius around Arsenic atom.
In total only one binding site of Arsenic was determined in the X-Ray Structure of the Ligand-Binding Core of GLUA2 in Complex with (S)-7-Hpca at 2.5 A Resolution, PDB code: 3pd8:
In total only one binding site of Arsenic was determined in the X-Ray Structure of the Ligand-Binding Core of GLUA2 in Complex with (S)-7-Hpca at 2.5 A Resolution, PDB code: 3pd8:
Arsenic binding site 1 out of 1 in 3pd8
Go back to
Arsenic binding site 1 out
of 1 in the X-Ray Structure of the Ligand-Binding Core of GLUA2 in Complex with (S)-7-Hpca at 2.5 A Resolution
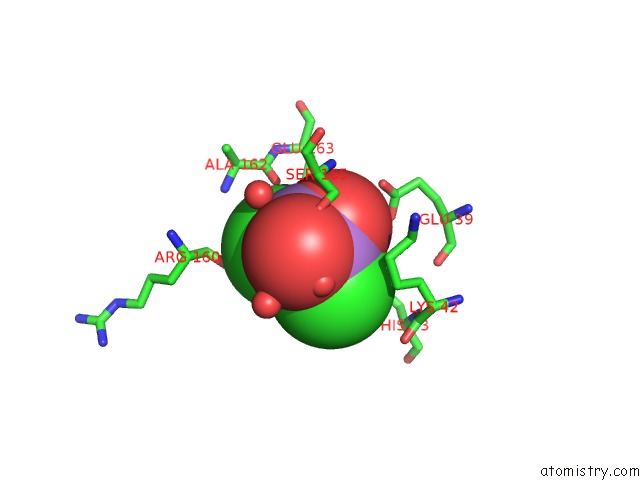
Mono view
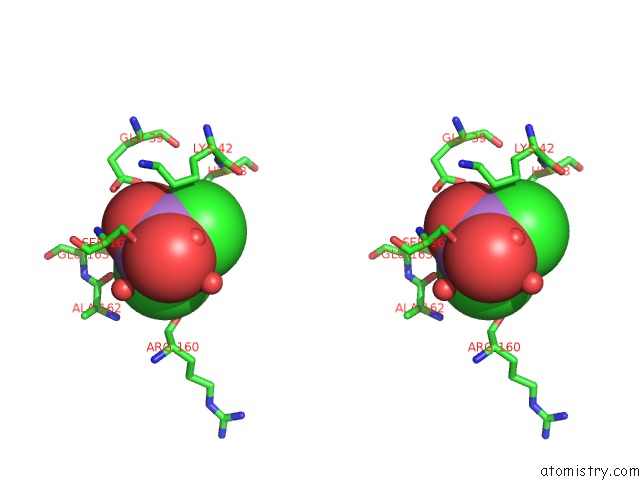
Stereo pair view

Mono view

Stereo pair view
A full contact list of Arsenic with other atoms in the As binding
site number 1 of X-Ray Structure of the Ligand-Binding Core of GLUA2 in Complex with (S)-7-Hpca at 2.5 A Resolution within 5.0Å range:
|
Reference:
K.Frydenvang,
D.S.Pickering,
J.R.Greenwood,
N.Krogsgaard-Larsen,
L.Brehm,
B.Nielsen,
S.B.Vogensen,
H.Hald,
J.S.Kastrup,
P.Krogsgaard-Larsen,
R.P.Clausen.
Biostructural and Pharmacological Studies of Bicyclic Analogues of the 3-Isoxazolol Glutamate Receptor Agonist Ibotenic Acid. J. Med. Chem. V. 53 8354 2010.
ISSN: ISSN 1520-4804
PubMed: 21067182
DOI: 10.1021/JM101218A
Page generated: Sun Jul 6 23:31:39 2025
ISSN: ISSN 1520-4804
PubMed: 21067182
DOI: 10.1021/JM101218A
Last articles
F in 7KYAF in 7KY5
F in 7KXW
F in 7KY9
F in 7KWA
F in 7KXT
F in 7KXC
F in 7KWV
F in 7KXL
F in 7KW4