Arsenic »
PDB 3n67-3smt »
3s2s »
Arsenic in PDB 3s2s: The Crystal Structure of Pyrazinamidase/Nicotinamidase From Streptococcus Mutans UA159
Enzymatic activity of The Crystal Structure of Pyrazinamidase/Nicotinamidase From Streptococcus Mutans UA159
All present enzymatic activity of The Crystal Structure of Pyrazinamidase/Nicotinamidase From Streptococcus Mutans UA159:
3.5.1.19;
3.5.1.19;
Protein crystallography data
The structure of The Crystal Structure of Pyrazinamidase/Nicotinamidase From Streptococcus Mutans UA159, PDB code: 3s2s
was solved by
X.-D.Su,
X.Liu,
H.Zhang,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 19.98 / 1.70 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 76.490, 80.120, 130.960, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 16 / 19.1 |
Other elements in 3s2s:
The structure of The Crystal Structure of Pyrazinamidase/Nicotinamidase From Streptococcus Mutans UA159 also contains other interesting chemical elements:
| Zinc | (Zn) | 4 atoms |
Arsenic Binding Sites:
The binding sites of Arsenic atom in the The Crystal Structure of Pyrazinamidase/Nicotinamidase From Streptococcus Mutans UA159
(pdb code 3s2s). This binding sites where shown within
5.0 Angstroms radius around Arsenic atom.
In total 4 binding sites of Arsenic where determined in the The Crystal Structure of Pyrazinamidase/Nicotinamidase From Streptococcus Mutans UA159, PDB code: 3s2s:
Jump to Arsenic binding site number: 1; 2; 3; 4;
In total 4 binding sites of Arsenic where determined in the The Crystal Structure of Pyrazinamidase/Nicotinamidase From Streptococcus Mutans UA159, PDB code: 3s2s:
Jump to Arsenic binding site number: 1; 2; 3; 4;
Arsenic binding site 1 out of 4 in 3s2s
Go back to
Arsenic binding site 1 out
of 4 in the The Crystal Structure of Pyrazinamidase/Nicotinamidase From Streptococcus Mutans UA159
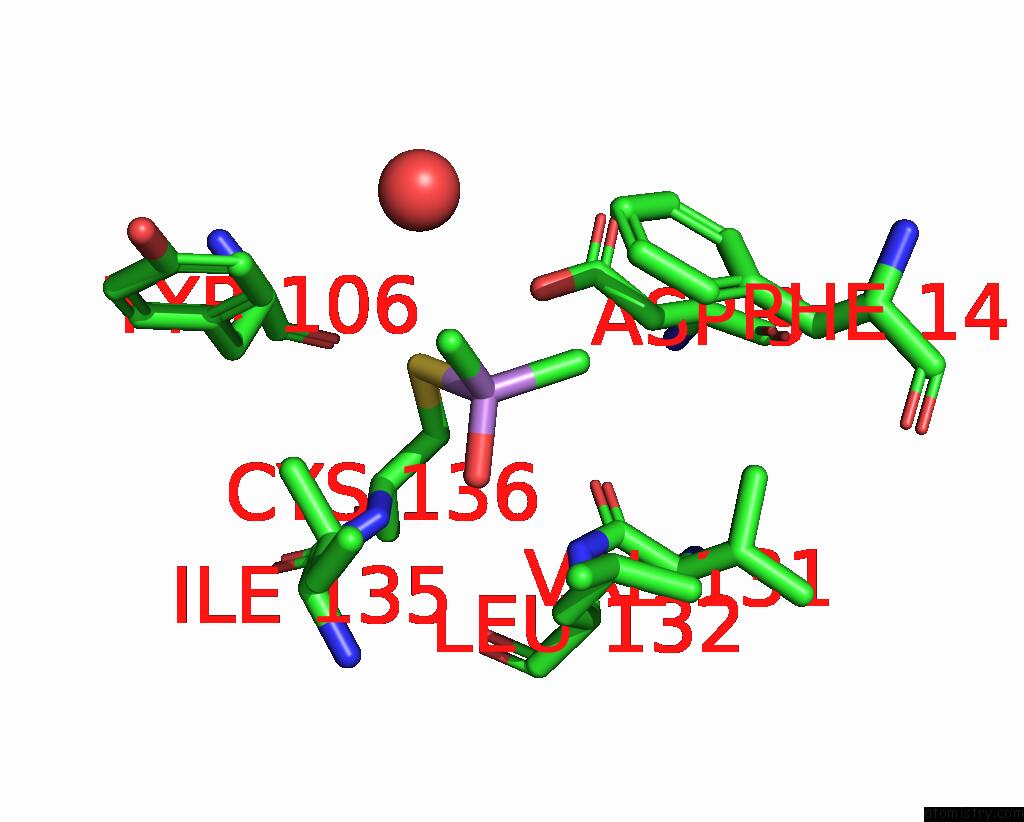
Mono view
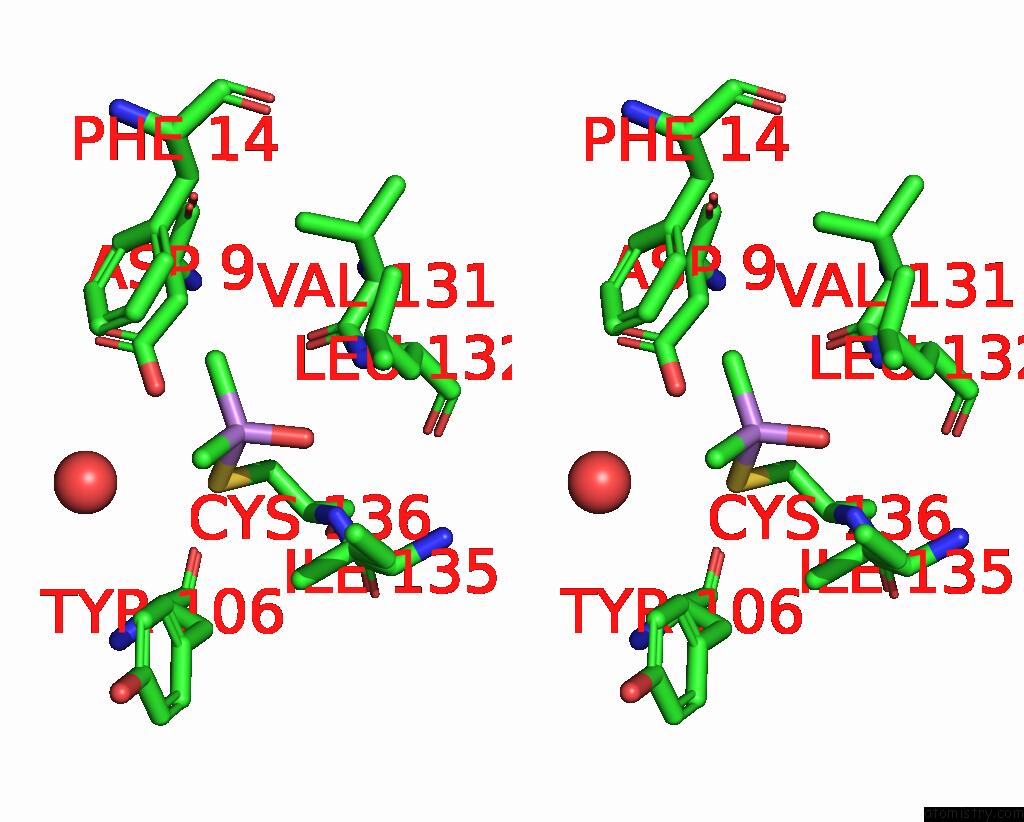
Stereo pair view

Mono view

Stereo pair view
A full contact list of Arsenic with other atoms in the As binding
site number 1 of The Crystal Structure of Pyrazinamidase/Nicotinamidase From Streptococcus Mutans UA159 within 5.0Å range:
|
Arsenic binding site 2 out of 4 in 3s2s
Go back to
Arsenic binding site 2 out
of 4 in the The Crystal Structure of Pyrazinamidase/Nicotinamidase From Streptococcus Mutans UA159
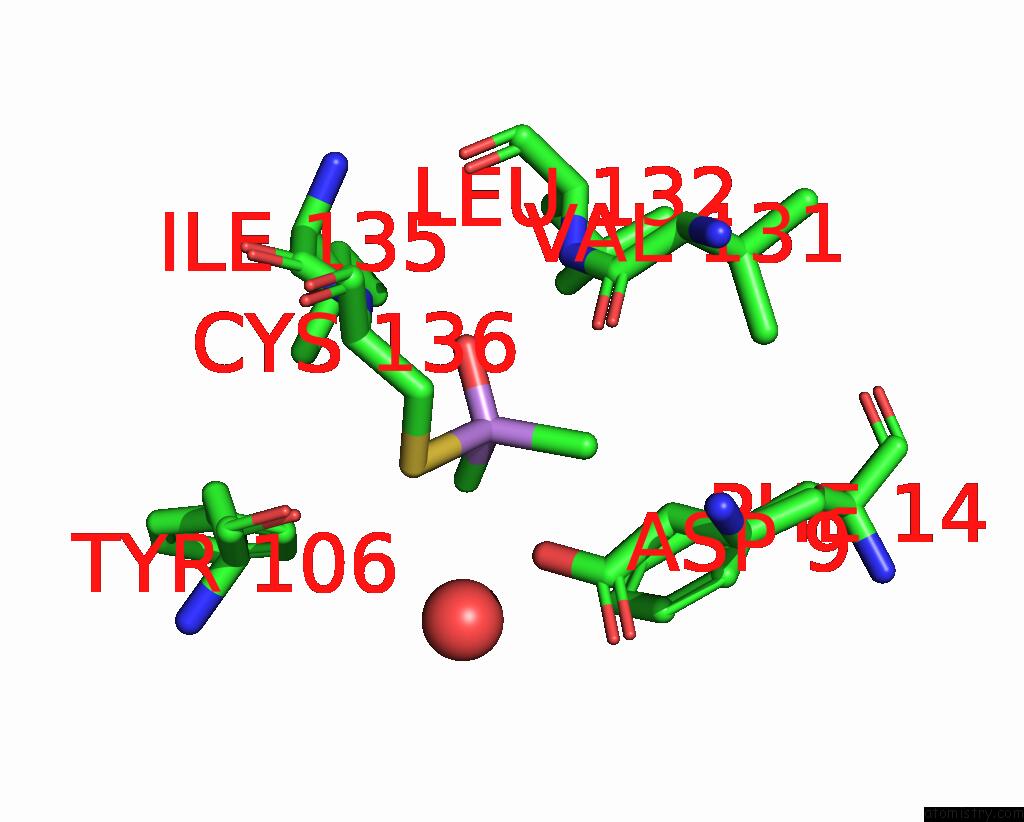
Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Arsenic with other atoms in the As binding
site number 2 of The Crystal Structure of Pyrazinamidase/Nicotinamidase From Streptococcus Mutans UA159 within 5.0Å range:
|
Arsenic binding site 3 out of 4 in 3s2s
Go back to
Arsenic binding site 3 out
of 4 in the The Crystal Structure of Pyrazinamidase/Nicotinamidase From Streptococcus Mutans UA159
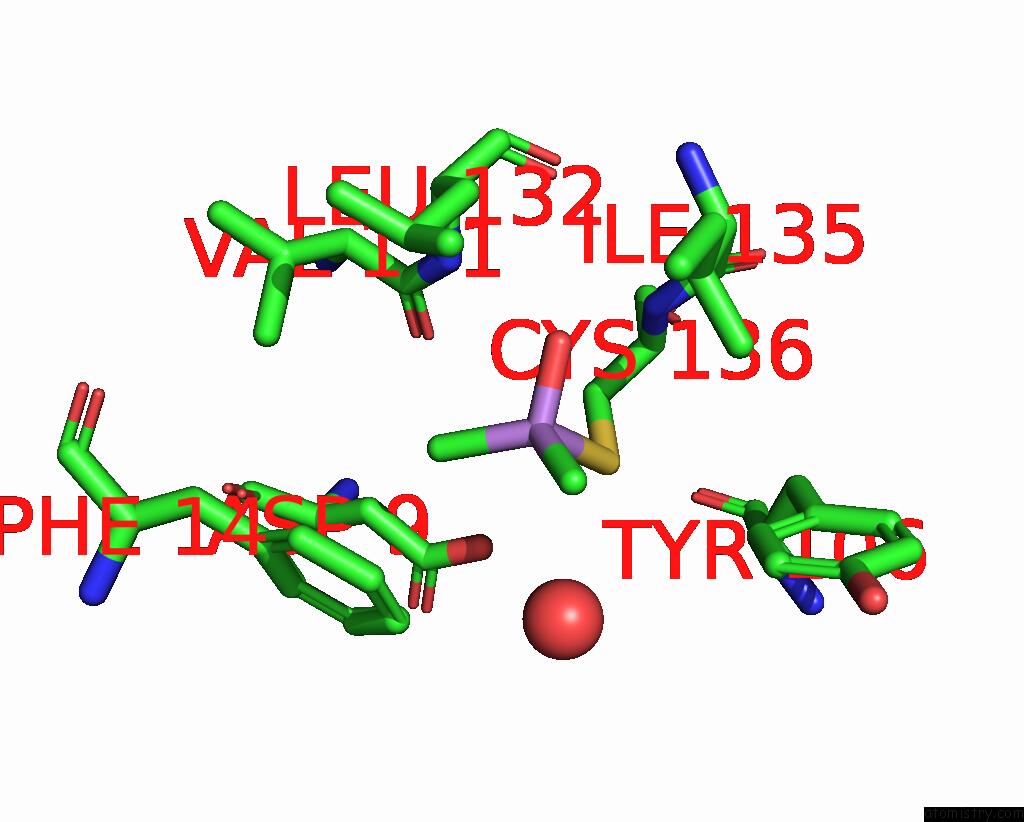
Mono view
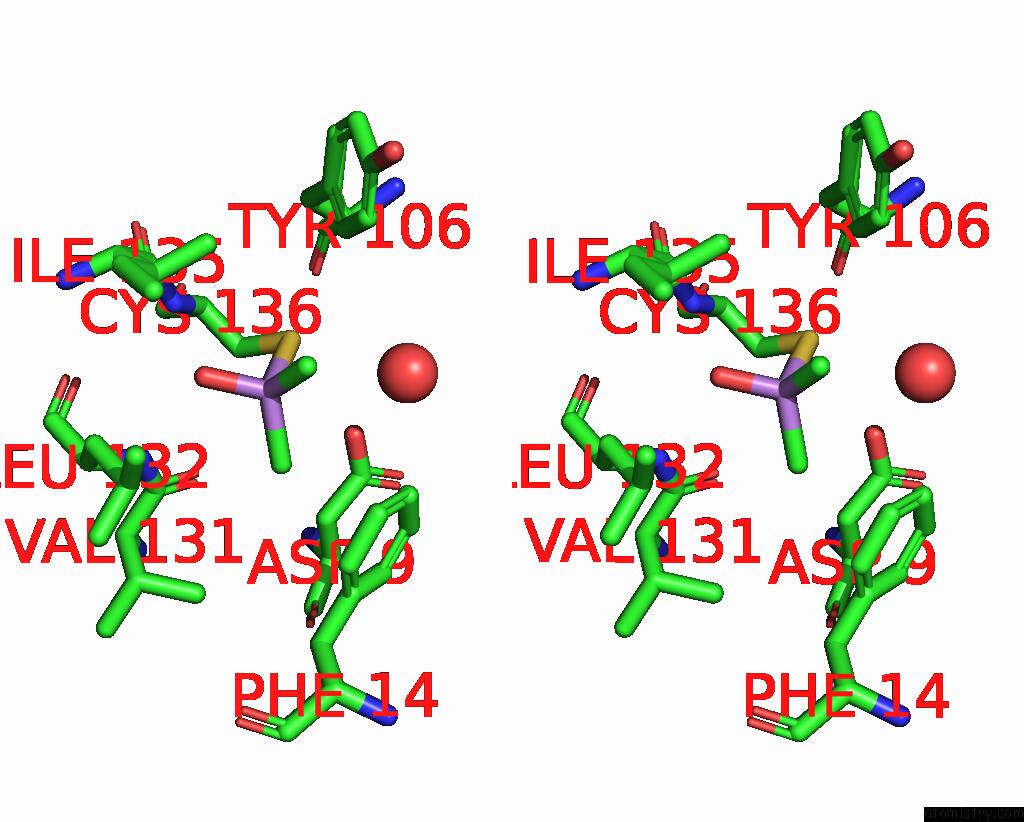
Stereo pair view

Mono view

Stereo pair view
A full contact list of Arsenic with other atoms in the As binding
site number 3 of The Crystal Structure of Pyrazinamidase/Nicotinamidase From Streptococcus Mutans UA159 within 5.0Å range:
|
Arsenic binding site 4 out of 4 in 3s2s
Go back to
Arsenic binding site 4 out
of 4 in the The Crystal Structure of Pyrazinamidase/Nicotinamidase From Streptococcus Mutans UA159
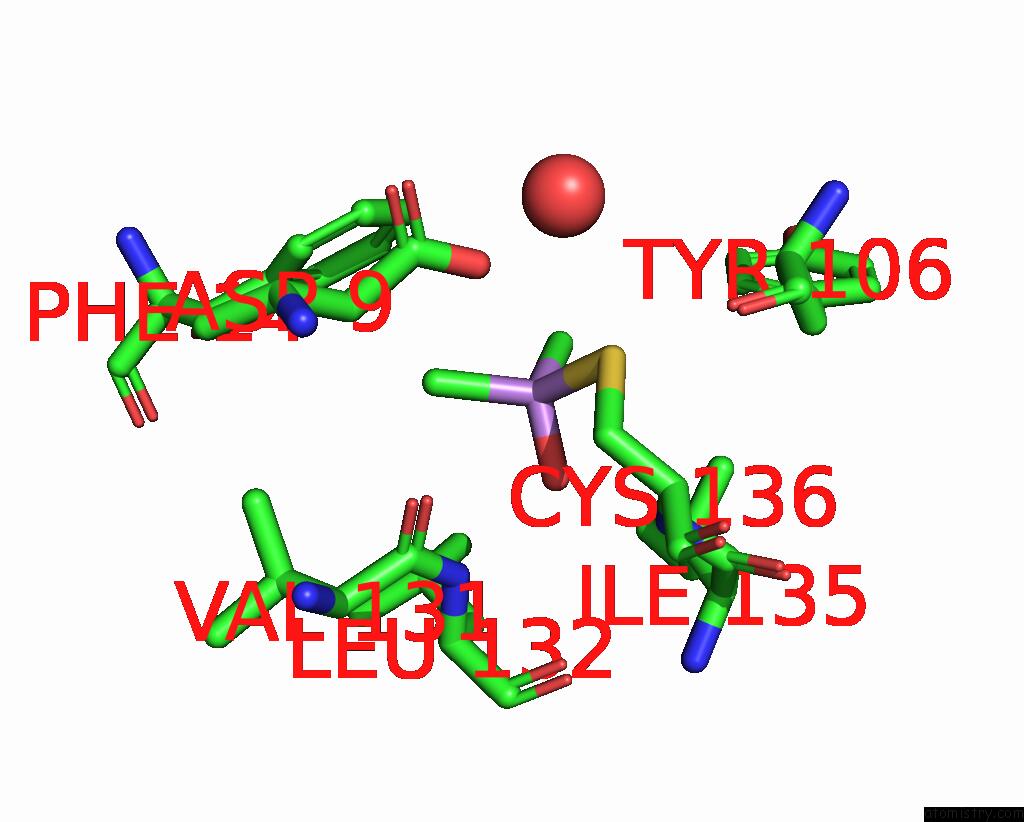
Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Arsenic with other atoms in the As binding
site number 4 of The Crystal Structure of Pyrazinamidase/Nicotinamidase From Streptococcus Mutans UA159 within 5.0Å range:
|
Reference:
X.Liu,
H.Zhang,
X.J.Wang,
L.F.Li,
X.-D.Su.
Get Phases From Arsenic Anomalous Scattering: De Novo Sad Phasing of Two Protein Structures Crystallized in Cacodylate Buffer Plos One V. 6 24227 2011.
ISSN: ESSN 1932-6203
PubMed: 21912678
DOI: 10.1371/JOURNAL.PONE.0024227
Page generated: Sun Jul 6 23:33:00 2025
ISSN: ESSN 1932-6203
PubMed: 21912678
DOI: 10.1371/JOURNAL.PONE.0024227
Last articles
F in 7G9KF in 7G9P
F in 7G7Z
F in 7G9N
F in 7G94
F in 7G9M
F in 7G8J
F in 7G8T
F in 7G8I
F in 7G8F